Back
BackProblem 16b
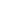
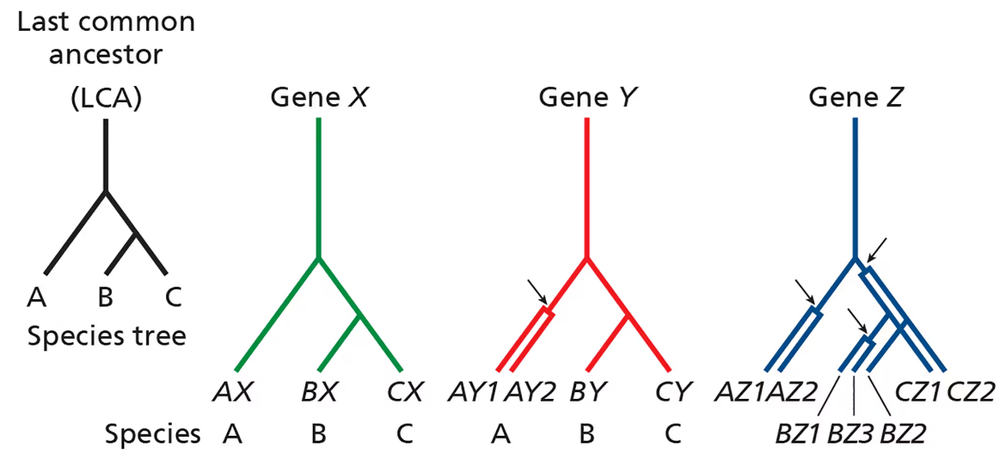
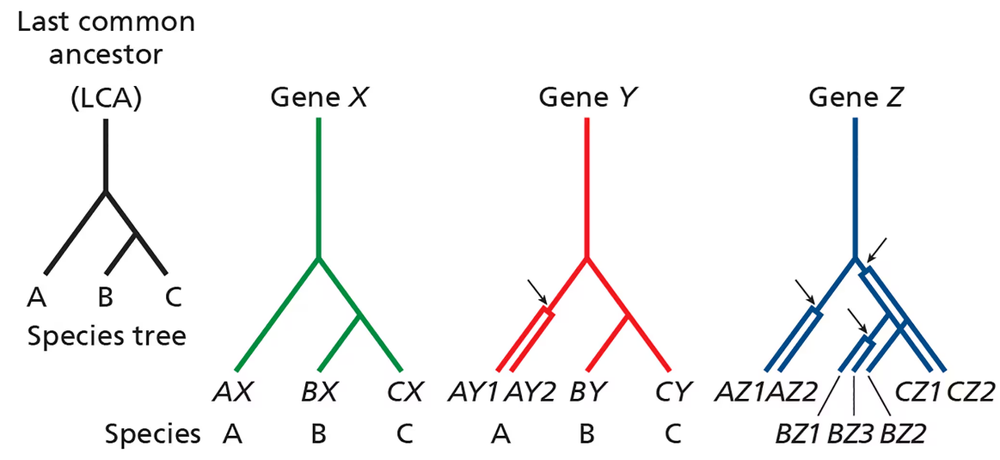
Consider the phylogenetic trees below pertaining to three related species (A, B, and C) that share a common ancestor (last common ancestor, or LCA). The lineage leading to species A diverges before the divergence of species B and C.
For gene Y, a gene duplication occurred in the lineage leading to A after it diverged from that, leading to B and C. Are genes AY1 and AY2 orthologous or paralogous? Are genes AY1 and BY orthologous or paralogous? Are genes BY and CY orthologous or paralogous?
Problem 16c
Consider the phylogenetic trees below pertaining to three related species (A, B, and C) that share a common ancestor (last common ancestor, or LCA). The lineage leading to species A diverges before the divergence of species B and C.
For gene Z, gene duplications have occurred in all species. Define orthology and paralogy relationships for the different Z genes.
Problem 17
You have isolated a gene that is important for the production of milk and wish to study its regulation. You examine the genomes of human, mouse, dog, chicken, pufferfish, and yeast and note that all genomes except yeast have an orthologous gene.
What does the existence of orthologous genes in chicken and pufferfish tell you about the function of this gene?Problem 17a
You have isolated a gene that is important for the production of milk and wish to study its regulation. You examine the genomes of human, mouse, dog, chicken, pufferfish, and yeast and note that all genomes except yeast have an orthologous gene.
How would you identify the regulatory elements important for the expression of your isolated gene in mammary glands?
Problem 18
When the human genome is examined, the chromosomes appear to have undergone only minimal rearrangement in the 100 million years since the last common ancestor of eutherian mammals. However, when individual humans are examined or when the human genome is compared with that of chimpanzees, a large number of small indels and SNPs can be detected. How are these observations reconciled?
Problem 19
Symbiodinium minutum is a dinoflagellate with a genome size that encodes more than 40,000 protein-coding genes. In contrast, the genome of Plasmodium falciparum has only a little more than 5000 protein-coding genes. Both Symbiodinium and Plasmodium are members of the Alveolate lineage of eukaryotes. What might be the cause of such a wide variation in their genome sizes?
Problem 20
Substantial fractions of the genomes of many plants consist of segmental duplications; for example, approximately 40% of genes in the Arabidopsis genome are duplicated. How might you approach the functional characterization of such genes using reverse genetics?
Problem 21
A modification of the two-hybrid system, called the one-hybrid system, is used for identifying proteins that can bind specific DNA sequences. In this method, the DNA sequence to be tested, the bait, is fused to a TATA box to drive expression of a reporter gene. The reporter gene is often chosen to complement a mutant phenotype; for example, a HIS gene may be used in a his⁻ mutant yeast strain. A cDNA library is constructed with the cDNA sequences translationally fused to the GAL4 activation domain and transformed into this yeast strain. Diagram how trans-acting proteins that bind to cis-acting regulatory sequences can be identified using a one-hybrid screen.
Problem 22a
A substantial fraction of almost every genome sequenced consists of genes that have no known function and that do not have sequence similarity to any genes with known function. Describe two approaches to ascertaining the biological role of these genes in S. cerevisiae.
Problem 22b
A substantial fraction of almost every genome sequenced consists of genes that have no known function and that do not have sequence similarity to any genes with known function. How would your approach change if the genes of unknown function were in the human genome?
Problem 23
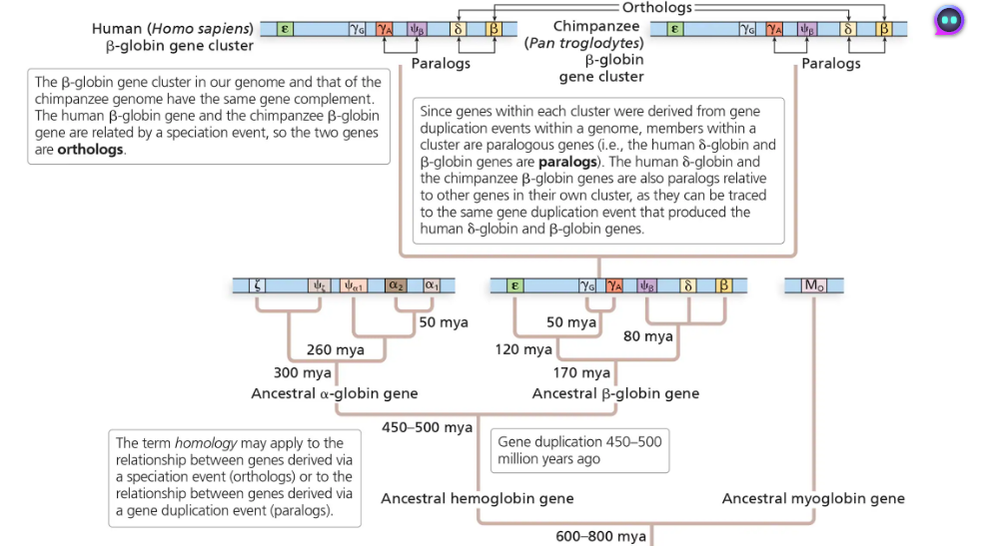
In the globin gene family (shown in the below diagram), which pair of genes would exhibit a higher level of sequence similarity, the human δ-globin and human β-globin genes or the human β-globin and chimpanzee β-globin genes? Can you explain your answer in terms of the timing of gene duplications?
Problem 24
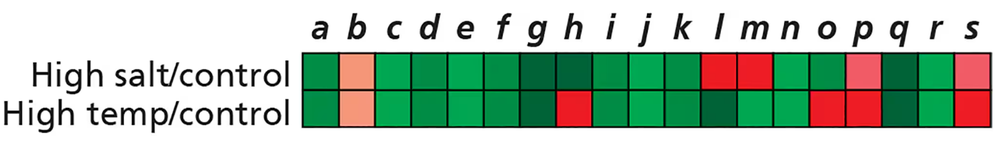
You are studying similarities and differences in how organisms respond to high salt concentrations and high temperatures. You begin your investigation by using microarrays to compare gene expression patterns of S. cerevisiae in normal growth conditions, in high salt concentrations, and at high temperatures. The results are shown here, with the values of red and green representing the extent of increase and decrease, respectively, of expression for genes a–s in the experimental conditions versus the control (normal growth) conditions. What is the first step you will take to analyze your data?
- In conducting the study described in Problem 24, you have noted that a set of S. cerevisiae genes are repressed when yeast are grown under high-salt conditions. How might you determine whether this set of genes is regulated by a common transcription factor?
Problem 25
Problem 25b
In conducting the study described in Problem 24, you have noted that a set of S. cerevisiae genes are repressed when yeast are grown under high-salt conditions. How might you approach this question if genome sequences for the related Saccharomyces species S. paradoxus, S. mikatae, and S. bayanus were also available?
Problem 26
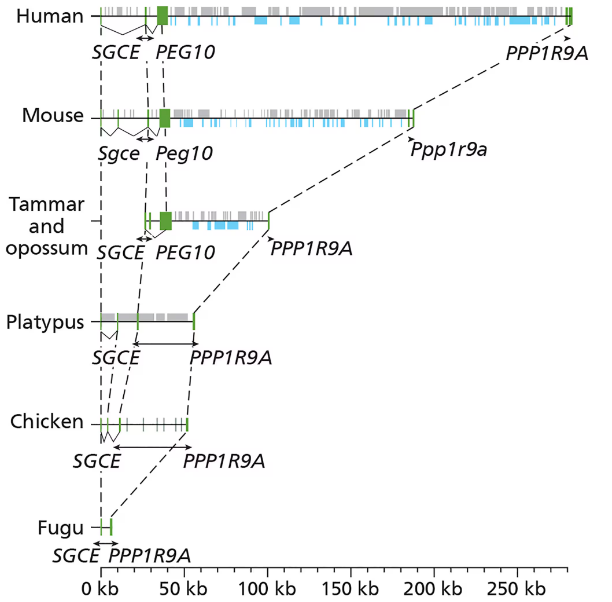
PEG10 (paternally expressed gene 10) is a paternally expressed gene (meaning only the paternal allele is expressed) that has an essential role in the formation of the placenta of the mouse. In the mouse genome, the PEG10 gene is flanked by the SGCE and PPP1R9A genes. To study the origin of PEG10, you examine syntenic regions spanning the SGCE and PPP1R9A loci in the genomes of several vertebrates, and you note that the PEG10 gene is present in the genomes of placental and marsupial mammals but not in the platypus, chicken, or fugu genomes.
The green bars in the figure indicate the exons of each gene. The gray bars represent LINEs and SINEs, and the blue bars represent long terminal repeat (LTR) elements of retrotransposons. Solid black diagonal lines link introns, and dashed black lines connect orthologous exons. Arrowheads indicate the direction of transcription.
Using the predicted protein sequence of PEG10, you perform a tblastn search for homologous genes and find that the most similar sequences are in a class of retrotransposons (the sushi-ichi retrotransposons). Propose an evolutionary scenario for the origin of the PEG10 gene, and relate its origin to its biological function.
Problem 27
What is the difference between biochemical and biological function?
Problem 28
Using the two-hybrid system to detect interactions between proteins, you obtained the following results: A clone encoding gene A gave positive results with clones B and C; clone B gave positive results with clones A, D, and E but not C; and clone E gave positive results only with clone B. Another clone F gave positive results with clone G but not with any of A–E. Can you explain these results? To follow up your two-hybrid results, you isolate null loss-of-function mutations in each of the genes A–G. Mutants of genes A, B, C, D, and E grow at only 80% of the rate of the wild type, whereas mutants of genes F and G are phenotypically indistinguishable from the wild type. You construct several double-mutant strains: The ab, ac, ad, and ae double mutants all grow at about 80% of the rate of the wild type, but af and ag double mutants exhibit lethality. Explain these results. How do the two-hybrid system and genetic interaction results complement one another? Can you reconcile your two-hybrid system and genetic interaction results in a single model?
Problem 29
Describe at least two mechanisms by which duplicate genes arise. What are the possible fates of duplicate genes? Does the mode of duplication affect possible fates?
Problem 33
Describe how enhancer screens can be used to uncover genetic redundancy.